|
9/20/2023 0 Comments Pet28a vector snapgene 
The higher the percentage of agarose, the smaller the effective range of separation will be. Depending on the size of the DNA fragments you wish to analyze, more of less agarose can be incorporated into a DNA gel. Here we will use a 500 bp ladder that includes bands of DNA that are 500 bp, 1000 bp, 1500 bp, 2000 bp, and so on. The size (in bp) of each separated DNA fragment is determined by comparison to a DNA ladder that has a band at given intervals of bp. The smaller molecule travel faster than the larger molecules. Due to the negatively charged phosphate backbone of DNA, DNA molecules migrate from the negatively-charged to the positively-charged end of the gel. 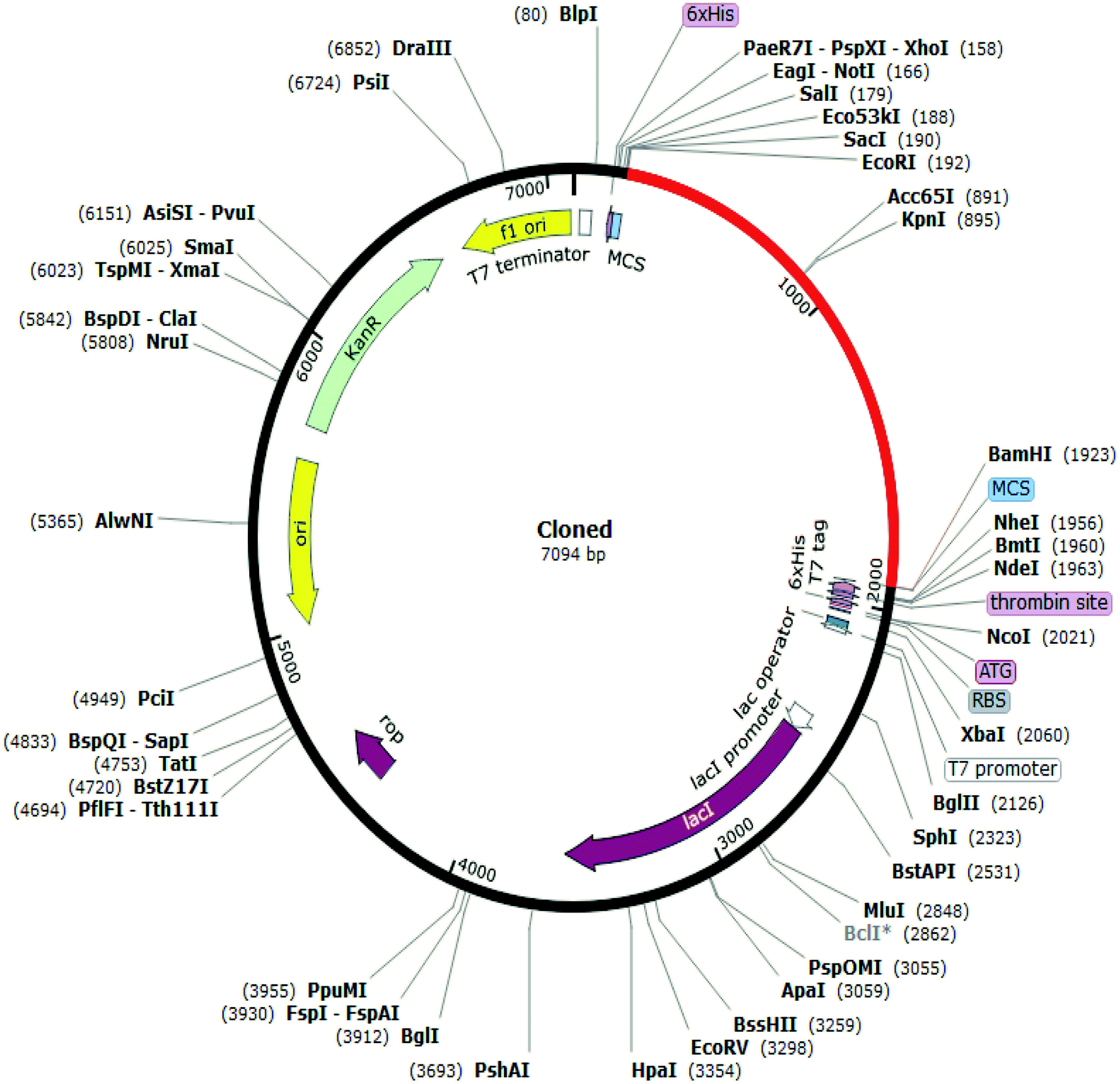
When the digestions are complete, remove the tubes from the incubator and add 4 μL of 6X nucleic acid loading buffer to each of the reactions.ĭNA electrophoresis enables size-based DNA separation. Meanwhile, prepare an agarose gel (described below) to analyze the digested DNA by electrophoresis. Incubate the digestions for 1 hour in the 37 ✬ incubator. Mix the reactions gently by pipetting, close the tubes, and centrifuge for several seconds in a bench-top microcentrifuge. Mix by pipetting, then add: Nde1, 10 units/ μL Mix by pipetting, then add: Xho1, 10 units/ μL In each of four sterile 0.65 mL tubes, add the following components in the order listed, where x = the volume of miniprep elution added (in μL) to give 1 μg of DNA: Set up analytical-scale digests of your plasmid DNA using 1 μg of DNA and final reaction volumes of 20 μL each. A sample gel is provided below.ġ.) DNA digestion with Xho1 and Nde1 restriction enzymes Since coiled plasmid DNA travels further by electrophoresis than equivalent linearized DNA, these reactions should result in bands with an apparent size slightly larger than the uncut plasmid. The oneenzyme reactions should linearize the plasmid DNA without excising any segment. By gel analysis of the four reactions, you will expect to see a band for the no-enzyme “mock” reaction at approximately 6,138 bp (consistent with the size of the insert plus vector. As experimental controls, you will simultaneously prepare a no-enzyme “mock” digest and digests containing only one of the two restriction enzymes required to excise the ABL insert. As denoted in the vector map, the pET28a vector (minus the segment between the Nde1 and Xho1 sites) is 5,289 bp, and the ABL insert should be 849 bp. Today you will excise the insert by digestion of the isolated plamid DNA with Xho1 and Nde and check the size of the digestion products on an agarose gel to confirm that your vector and insert are the expected size in base pairs (bp). Since the gene encoding the Abl kinase domain is positioned in the pET-28a vector between Xho1 and an Nde1 recognition sites, treatment of the plasmid with those two restriction enzymes should result in excision of the ABL insert. In Session 2, you isolated a DNA plasmid (the pET-28a vector) containing an insert encoding residues 229-511 of the Abl kinase domain. In DNA engineering, restriction enzymes are used to cut open plasmids for the ligation of a complementarily-cut DNA insert into the plasmid for protein expression. Restriction enzymes (RE’s) are enzymes that selectively cleave double stranded DNA based on recognition of specific, short DNA sequences called recognition sites. In session 4, you will also prepare a standard curve for future protein quantification. In preparation for purifying the H396P Abl kinase domain, you will prepare all the necessary buffers for the lysis and purification. You will then design primers for subsequent site-directed mutagenesis. \)ĭuring sessions 3 and 4 you will verify that the plasmid DNA you isolated contains a construct of the expected size for the Abl kinase domain. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
